PRTools, elements, operations, user commands, introductory examples, advanced examples
Datasets
On this page the dataset is introduced. It is one of the key elements of PRTools
Dataset background
The dataset is a PRTools variable type in which the vector representation of a set of objects is stored together with object class labels, class priors, and other useful annotation of the objects, the measurements or the dataset in its entirety. The vector elements traditionally correspond to features, but may also refer to pixel values in an image, spectral intensities for a set of wavelengths, amplitudes of a time signal at given points in time, dissimilarities with a fixed set of other objects, etcetera. Essential is that every object the same set measurement values are available.
The core of a dataset is a data matrix of m rows (corresponding to the objects) and k columns (corresponding to the measurements). The columns are usually referred to as features but they may have another background as in the above examples. A dataset is hereby an enriched version of a data matrix.It has a size of m*k and many of the Matlab operations for matrices apply. In addition PRTools adds specific dataset operations and, most important, the large set of routines for dimension reduction, classification, evaluation and visualization know how to handle datasets. This avoids that users have to manipulate additional parameters and annotation themselves.
Dataset definition
The class of dataset variables is called prdataset. In the past it was just, natually, dataset, but this was changed in PRTools5 to avoid clashes with other toolboxes.
The prdataset constructor looks like
a = prdataset(data,labels)
datais an array of size[m,k]storing thek-dimensional vector representations ofmobjects.labelscontains themobject labels. The following types are supported:
– numeric: a columns vector ofkintegers (users are discouraged to use this type).
– strings: a character array okkstrings.
– cells: a 1-dimensional vector ofkcells, each containing a string.
The two items data and labels are essential for the operation of PRTools. If data is neglected (data = []) an empty dataset is defined. If labels is not supplied the objects remain unlabeled. In case just a part of the objects have no labels the corresponding entry of labels should contain a NaN for numeric labels or an empty string ('') in case of string labels.
In order to speed up the handling of labels they are stored in a (character) array of class names (called lablist). Instead of dealing with the possibly long names, PRTools just uses for every object an index (called nlab) into lablist. For more information see the description of the prdataset structure and the commands for storing information into a dataset and retrieving it.
There are three types of labels supported for datasets:
- crisp. These are the traditional nominal class labels, either integer numbers or strings (class names). Object can belong to just a single class.
- soft. These labels are real values interval [0,1]. For every class a value has to be specified, not necessarily summing to one. They may be interpreted as fuzzy memberships, as class confidences or as posterior class probabilities.
- targets. These are numeric values on the interval (-infinity,infinity). Every object has for every class a target. This facility enables the construction of routines for multiple regression within the
PRToolsframework. There are just a few routines that use this facility.
Dataset overload
Although datasets may carry various kinds of additional information, they can be treated like 2-dimensional matrices in many common Matlab constructs. See the section dataset operations for examples. The resulting datasets always have, as far as applicable, the same fields as the original dataset. Where needed they are adapted (e.g. sizes and labels) to assure that the result is a consistent dataset.
Dataset structure
Various items can be stored in a dataset. A full list can be found be converting a dataset variable into a structure.
data = rand(6,2);
labels = [1 1 1, 2 2 2]';
a = prdataset(data,labels)
% 6 by 2 dataset with 2 classes: [3 3]
struct(a)
%
% data: [6x2 double]
% lablist: {2x4 cell}
% nlab: [6x1 double]
% labtype: 'crisp'
% targets: []
% featlab: []
% featdom: {}
% prior: []
% cost: []
% objsize: 6
% featsize: 2
% ident: [1x1 struct]
% version: {[1x1 struct] '03-Jan-2011 15:24:04'}
% name: []
% user: []
All fields have a corresponding set-command (e.g. setdata) to store it and a get-command (e.g. getdata) to retrieve it. Users are discouraged to use the ‘.’-constructs (e.g. a.files) as it will not guarantee consistency with other fields. In some cases not the exact fields are retrieved but some derived data. In the table more information is given.
| > The fields of the dataset structure | |
data |
This is the main field, storing the data as it is supplied by calling the dataset constructor or by setdata. The size of dataset, the number of objects (m) by the number of features (k) is derived from data. |
lablist |
This is a cell array that encodes the class names derived from the objects labels. Datasets can have multiple sets of labels for their objects of which just one is active, multi-labeling. The lablist field stores the necessary administration. The active set of labels can be retrieved by the commands classnames and getlablist. See also the following nlab item. |
nlab |
The labels supplied in the dataset definition are summarized by lablist and nlab. lablist contains the unique labels (class names) and nlab is an index vector for lablist. Its values range between 1 and the total number of classes (size of lablist). Entries in nlab for objects that are unlabeled are set to 0 (zero). The setnlab command should be treated with care as it changes the labeling of the dataset. |
labtype |
The labeling type (crisp, soft or targets, see above) is stored here. setlabtype changes the label type, but may also change nlab and lablist fields. |
featlab |
The feature labels are strings or numbers and are used by PRTools (if given) to annotate plots. |
featdom |
Here feature domains are stored. If these fields are set tests are performed whenever the values in the data field changes to check whether the new data is within the supplied domains. |
prior |
Classes in a dataset may have prior probabilities. These are used in density based classifiers, in error evaluation by testc and on some other places. If not set, the prior field is empty and when prior probabilities are needed the class frequencies in the dataset are taken. |
cost |
In this field a cost matrix can be stored for performance evaluation and procedures that explicitly minimize classification costs. Unless explicitly mentioned PRTools neglects this field. |
objsize |
The object size is the number of rows (objects) in the data field. It is retrieved by the size and getsize commands (getsize may return the number of classes as well). Although the routines getobjsize and setobjsize exist, users are discouraged to use them except in relation with image handling. |
featsize |
The feature size is the number of columns (features) in the data field. It is retrieved by the size and getsize commands (getsize retrieves the number of classes as well). Although the routines getfeatsize and setfeatsize exist, users are discouraged to use them except in relation with image handling. |
ident |
In subfields of the ident field various object identifiers can be stored. One field is always available: ident. Unless changed by the user it contains the object indices at creation of the dataset. Every ident subfield stores vectors or arrays of doubles, strings or cells with as many rows as there are objects in the dataset. |
version |
At creation of the dataset PRTools stores here its version and the date. |
name |
The user may supply a name here. It is displayed in the command window when a command returning a dataset is executed without a semicolon. The dataset name may also be used for annotating plots. |
user |
In this field the user can add and retrieve any additional annotation for the dataset in its entirety. |
When datasets are changed, e.g. by a transformation of the data, or by taking a subset of features or objects, all relevant information is copied, including the name and user field.
See the commands for the dataset definition and multi-labeling for more information on how to access the fields
Examples
Below some examples are given of dataset manipulations. Use is made of the PRTools command genlab(N) that generates a set of numeric labels, N(i) for class i. The command scatterd is similar but not identical to the Matlab command scatter and has thereby a similar, slightly different name.
% delete all figure
delfigs
% reset random seed for repeatability
% randreset(1)
% Generate in 2 dimensions 3 normally distributed classes
% of 20 objects for each class
a = prdataset(randn(60,2),genlab([20 20 20]))
% 60 by 2 dataset with 3 classes: [20 20 20]
% Give the features a name
a = setfeatlab(a,char('size','intensity'))
% 60 by 2 dataset with 3 classes: [20 20 20]
% Make the distributions of the classes different and plot them
a(1:20,:) = a(1:20,:)*0.5;
a(21:40,1) = a(21:40,1)+4;
a(41:60,2) = a(41:60,2)+4;
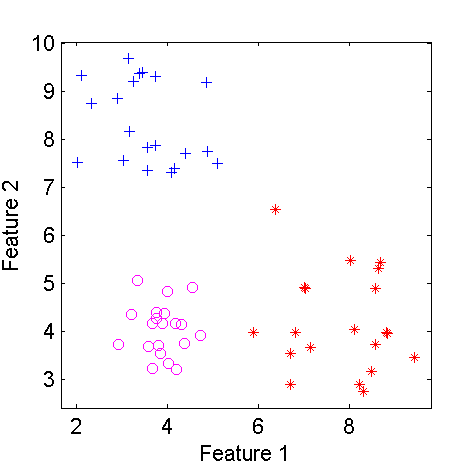
figure; scatterd(a)
% create a subset of the second class
b = a(21:40,:)
% 20 by 2 dataset with 3 classes: [0 20 0]
% add 4 to the second feature of this class
b(:,2) = b(:,2) + 4*ones(20,1)
% 20 by 2 dataset with 3 classes: [0 20 0]
% concatenate this set to the original dataset
c = [a;b]
% 80 by 2 dataset with 3 classes: [20 40 20]
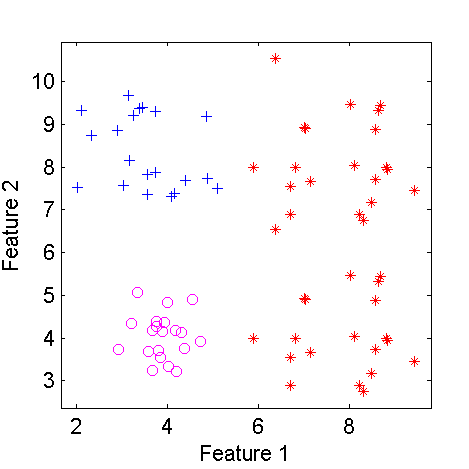
figure; scatterd(c);
showfigs
| Scatterplot of a three class two-dimensional dataset | The dataset after modifying class 2 |
For better annotation of the plot we may add some information on the dataset, the classes and features in some recognizable way, e.g.
c = setname(c,'Fruit dataset');
c = setlablist(c,char('apple','banana','cherry'));
c = setfeatlab(c,char('size','weight'));
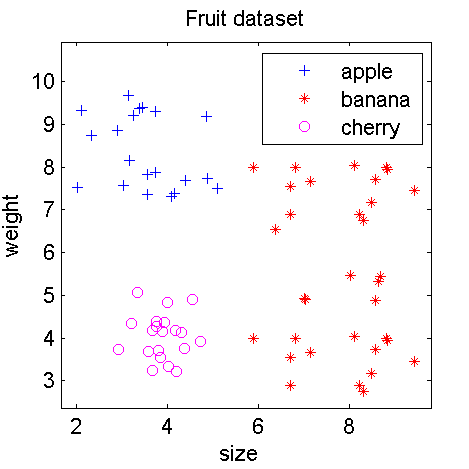
figure; scatterd(c)
The annotated dataset
elements:
datasets
datafiles
cells and doubles
mappings
classifiers
mapping types.
operations:
datasets
datafiles
cells and doubles
mappings
classifiers
stacked
parallel
sequential
dyadic.
user commands:
datasets
representation
classifiers
evaluation
clustering
examples
support routines.
introductory examples:
Introduction
Scatterplots
Datasets
Datafiles
Mappings
Classifiers
Evaluation
Learning curves
Feature curves
Dimension reduction
Combining classifiers
Dissimilarities.
advanced examples.